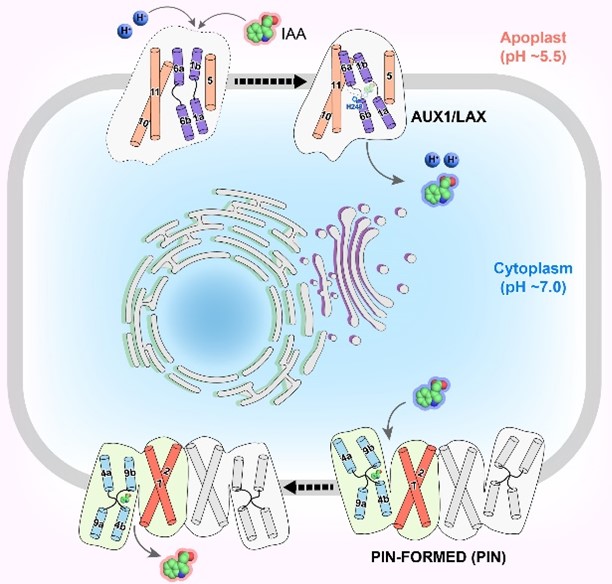
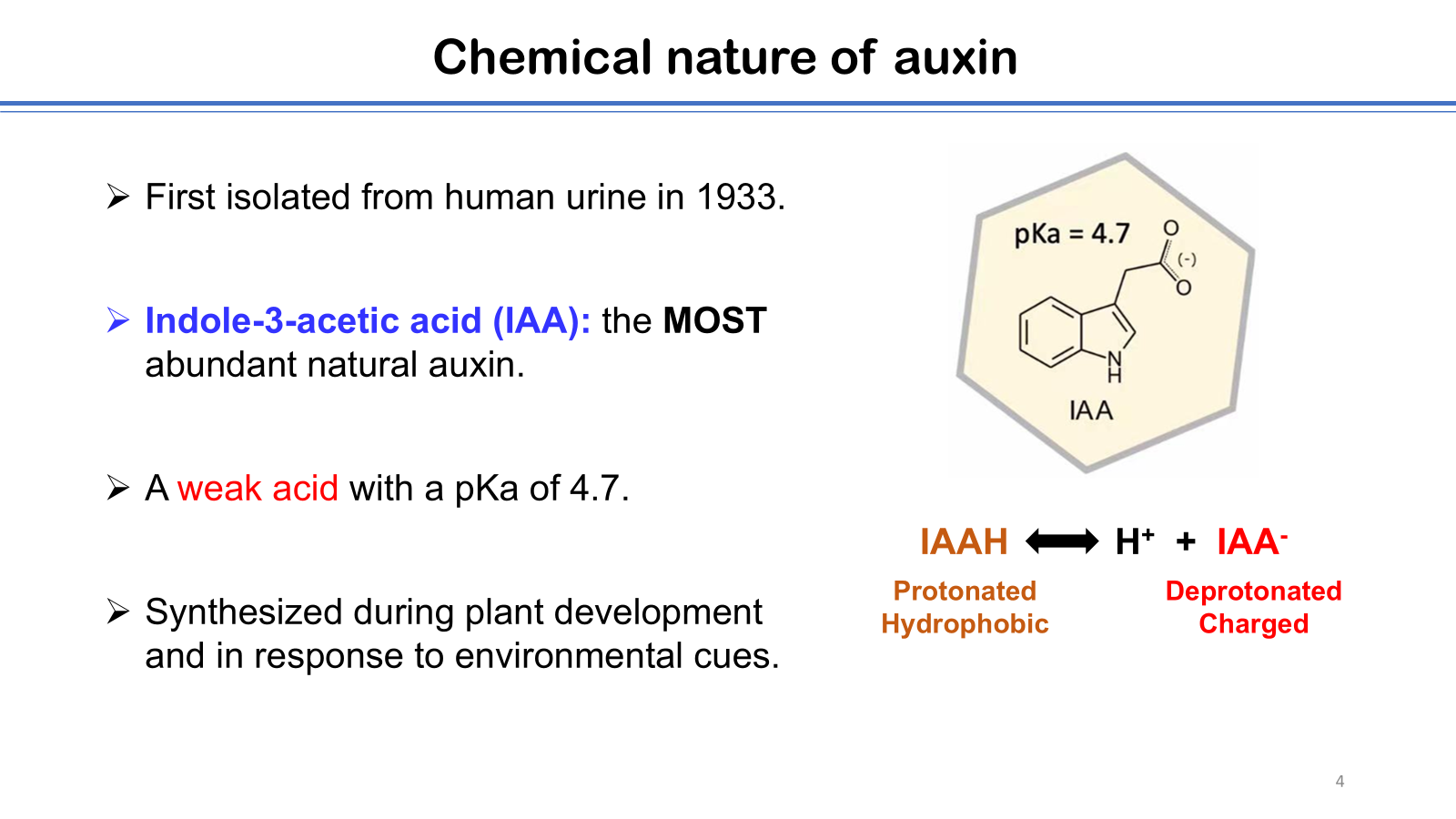
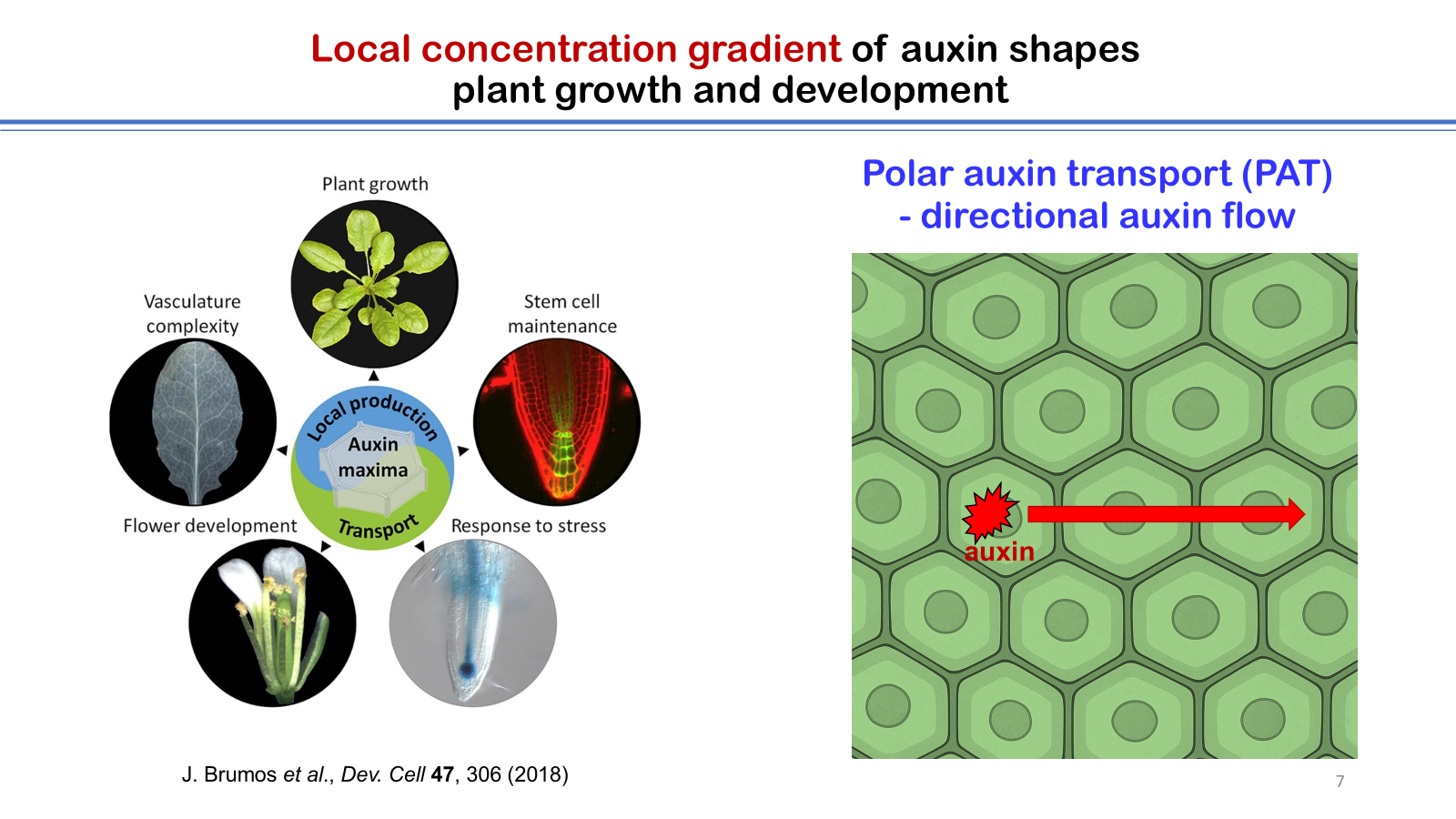
The author highlights his group’s research on the mechanistic characterizations of auxin exporters and importers in plants, with particular emphasis on the structure determination of these transporters using cryo-electron microscopy (cryo-EM). Auxin, the first plant hormone to be identified, is involved in nearly all aspects of plant growth, development, and responses to environmental cues. The biological activity of auxin depends on the establishment of local concentration gradients, which are generated through the process of polar auxin transport (PAT). Members of the PIN-FORMED (PIN) protein family and the AUXIN1/LIKE-AUXIN1 (AUX1/LAX) family are key components of PAT, mediating cellular auxin efflux and influx, respectively. Using cryo-EM single particle analysis, high-resolution structures of PIN1 and AUX1 were determined, revealing the overall architecture of each transporter as well as their modes of auxin and inhibitor binding. Biochemical analyses further demonstrated distinct transport mechanisms, showing that PIN1 functions as an auxin uniporter, whereas AUX1 acts as an auxin and proton symporter. Together, these findings advance our understanding of the molecular basis of PAT in plants, and provide a foundation for future agricultural applications targeting auxin transporters.
We acknowledge support from the National Natural Science Foundation of China (32322041, 32321001, and W2412029), the Scientific Research Innovation Capability Support Project for Young Faculty (ZYGXQNJSKYCXNLZCXM-B8 to L.S.), and Natural Science Foundation of Anhui Province (2408085JX005 to L.S.).
1. K.U. Torii, S. Hagihara, N. Uchida, and K. Takahashi, Harnessing synthetic chemistry to probe and hijack auxin signaling, New Phytol. 220, 417 (2018).
2. X. Cang, Q. Wang, L. Li, H. Han, and M. Chang, Auxin reimagined: transport and signaling patterns, J. Integr. Plant Biol. 68, 6 (2025).
3. R. Huang, J. Wang, M. Chang, W. Tang, Y. Yu, Y. Zhang, Y. Peng, Y. Wang, Y. Guo, T. Lu, Y. Cao, Y. Zhou, Q. Zhang, Y. Huang, A. Wu, L. Ren, M. Gallei, J. Dong, H. Chen, J. He, M. Wen, J. Friml, L. Sun, Y. Xiong, Z. Yang, and T. Xu, TMK–PIN1 drives a short self-organizing circuit for auxin export and signaling in Arabidopsis, Dev. Cell 61, 73 (2026).
4. J. Brumos, L.M. Robles, J. Yun, T.C. Vu, S. Jackson, J.M. Alonso, and A.N. Stepanova, Local auxin biosynthesis is a key regulator of plant development, Dev. Cell 47, 306 (2018).
5. R. Swarup and B. Péret, AUX/LAX family of auxin influx carriers—an overview, Front. Plant Sci. 3, 225 (2012).
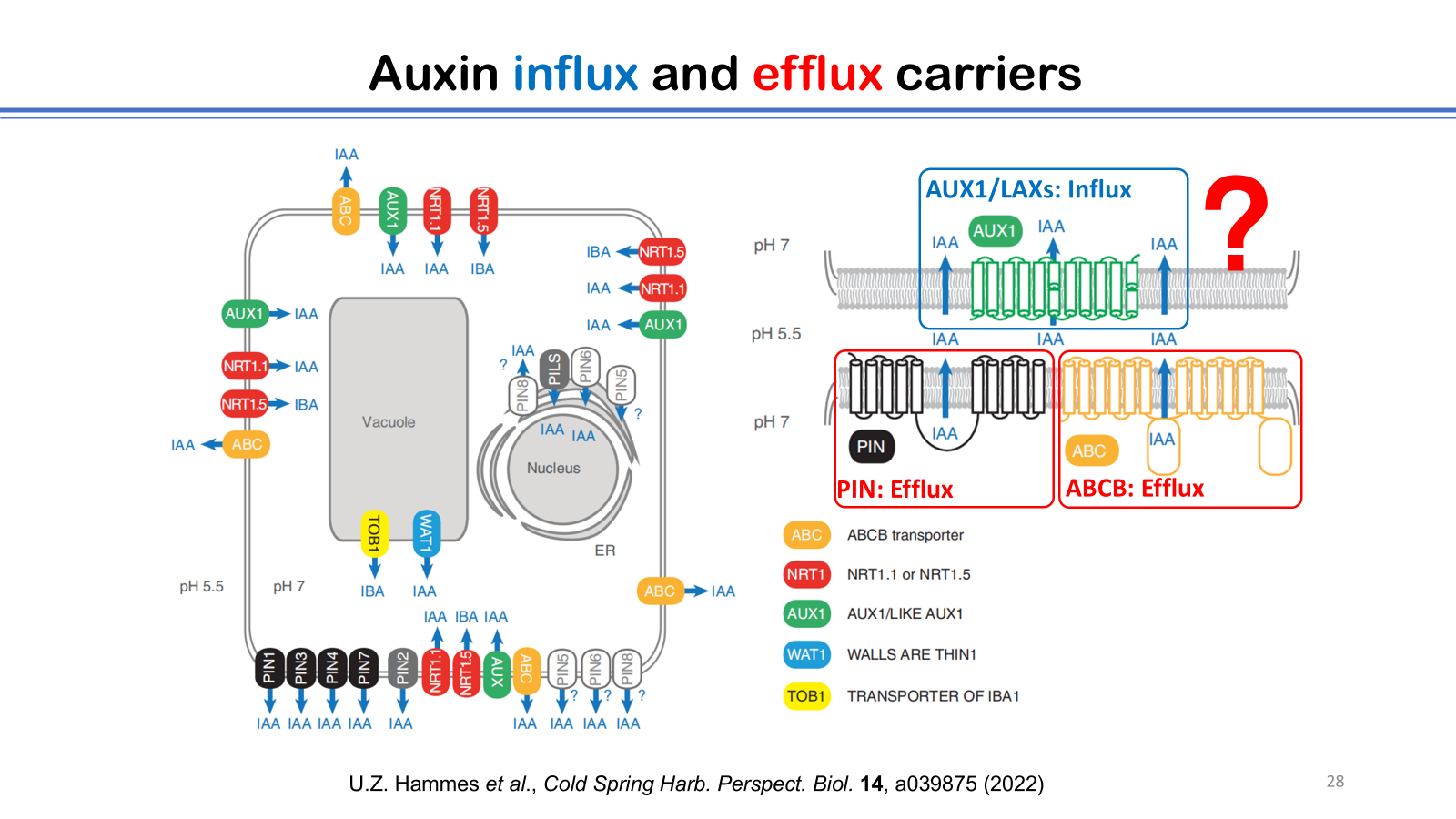
6. U.Z. Hammes, A.S. Murphy, and C. Schwechheimer, Auxin transporters—a biochemical view, Cold Spring Harb. Perspect. Biol. 14, a039875 (2022).
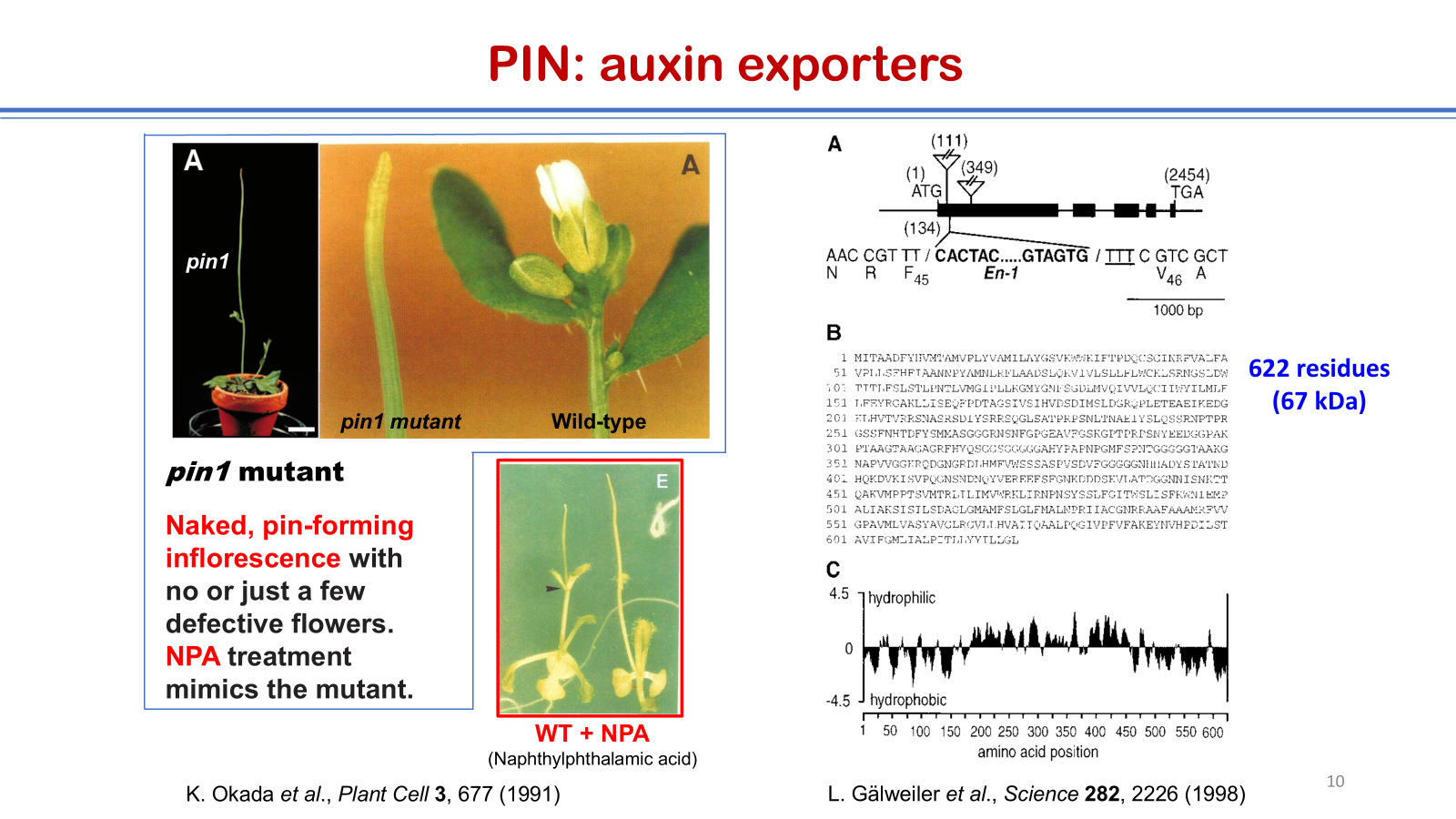
7. K. Okada, J. Ueda, M.K. Komaki, C.J. Bell, and Y. Shimura, Requirement of the auxin polar transport system in early stages of Arabidopsis floral bud formation, Plant Cell 3, 677 (1991).
8. L. Gälweiler, C. Guan, A. Müller, E. Wisman, K. Mendgen, A. Yephremov, and K. Palme, Regulation of polar auxin transport by AtPIN1 in Arabidopsis vascular tissue, Science 282, 2226 (1998).
9. M. Zwiewka, V. Bilanovičová, Y.W. Seifu, and T. Nodzyński, The nuts and bolts of PIN auxin efflux carriers, Front. Plant Sci. 10, 985 (2019).
10. M. Michniewicz, P.B. Brewer, and J. Friml, Polar auxin transport and asymmetric auxin distribution, Arabidopsis Book 2007, e0108 (2007).
11. A. Merk, A. Bartesaghi, S. Banerjee, V. Falconieri, P. Rao, M.I. Davis, R. Pragani, M.B. Boxer, L.A. Earl, J.L.S. Milne, and S. Subramaniam, Breaking cryo-EM resolution barriers to facilitate drug discovery, Cell 165, 1698 (2016).
12. Z. Yang, J. Xia, J. Hong, C. Zhang, H. Wei, W. Ying, C. Sun, L. Sun, Y. Mao, Y. Gao, S. Tan, J. Friml, D. Li, X. Liu, and L. Sun, Structural insights into auxin recognition and efflux by Arabidopsis PIN1, Nature 609, 611 (2022).
13. I. Zimmermann, P. Egloff, C.A.J. Hutter, F.M. Arnold, P. Stohler, N. Bocquet, M.N. Hug, S. Huber, M. Siegrist, L. Hetemann, J. Gera, S. Gmür, P. Spies, D. Gygax, E.R. Geertsma, R.J.P. Dawson, and M.A. Seeger, Synthetic single domain antibodies for the conformational trapping of membrane proteins, eLife 7, e34317 (2018).
14. K.L. Ung, L. Schulz, D.L. Stokes, U.Z. Hammes, and B.P. Pedersen, Substrate recognition and transport mechanism of the PIN-FORMED auxin exporters, Trends Biochem. Sci. 48, 937 (2023).
15. K.L. Ung, M. Winkler, L. Schulz, M. Kolb, D.P. Janacek, E. Dedic, D.L. Stokes, U.Z. Hammes, and B.P. Pedersen, Structures and mechanism of the plant PIN-FORMED auxin transporter, Nature 609, 605 (2022).
16. N. Su, A. Zhu, X. Tao, Z.J. Ding, S. Chang, F. Ye, Y. Zhang, C. Zhao, Q. Chen, J. Wang, C.Y. Zhou, Y. Guo, S. Jiao, S. Zhang, H. Wen, L. Ma, S. Ye, S.J. Zheng, F. Yang, S. Wu, and J. Guo, Structures and mechanisms of the Arabidopsis auxin transporter PIN3, Nature 609, 616 (2022).
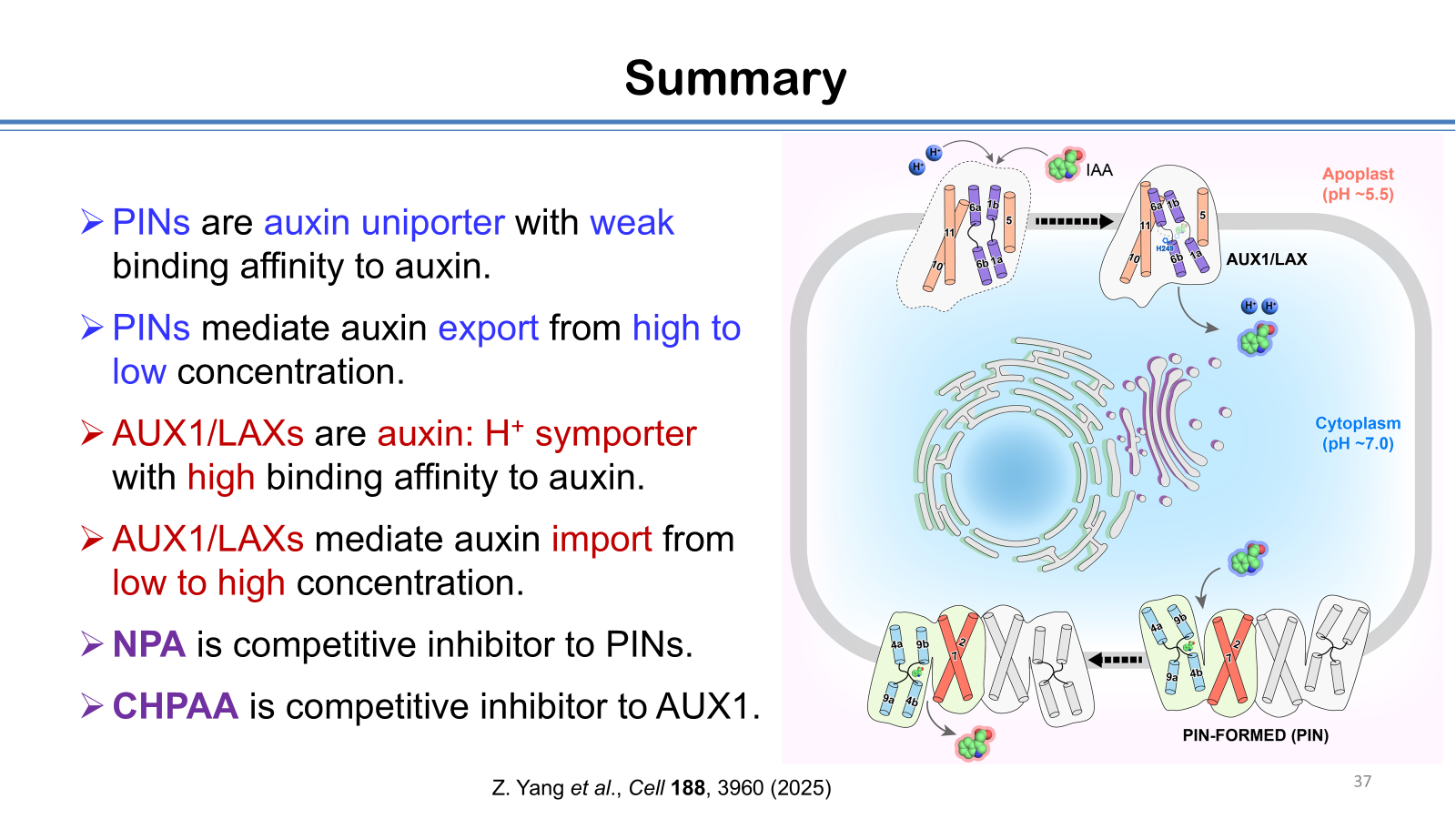
17. Z. Yang, H. Wei, Y. Gan, H. Liu, Y. Cao, H. An, X. Que, Y. Gao, L. Zhu, S. Tan, X. Liu, and L. Sun, Structural insights into auxin influx mediated by the Arabidopsis AUX1, Cell 188, 3960(2025).
18. W. Ying, Y. Wang, H. Wei, Y. Luo, Q. Ma, H. Zhu, H. Janssens, N. Vukašinović, M. Kvasnica, J.M. Winne, Y. Gao, S. Tan, J. Friml, X. Liu, E. Russinova, and L. Sun, Structure and function of the Arabidopsis ABC transporter ABCB19 in brassinosteroid export, Science 383, eadj4591 (2024).



